
 Data Structure
Data Structure Networking
Networking RDBMS
RDBMS Operating System
Operating System Java
Java MS Excel
MS Excel iOS
iOS HTML
HTML CSS
CSS Android
Android Python
Python C Programming
C Programming C++
C++ C#
C# MongoDB
MongoDB MySQL
MySQL Javascript
Javascript PHP
PHP
- Selected Reading
- UPSC IAS Exams Notes
- Developer's Best Practices
- Questions and Answers
- Effective Resume Writing
- HR Interview Questions
- Computer Glossary
- Who is Who
How to join points on a scatterplot with smooth lines in R using plot function?
It is very difficult to join points on a scatterplot with smooth lines if the scatteredness is high but we might want to look at the smoothness that cannot be understood by just looking at the points. It is also helpful to understand whether the model is linear or not. We can do this by plotting the model with loess using plot function.
Example
Consider the below data −
> set.seed(3) > x<-sample(1:100,10,replace=TRUE) > y<-rpois(10,100)
Using loess to create the smooth lines −
> Model <- loess(y~x) > summary(Model) Call: loess(formula = y ~ x) Number of Observations: 10 Equivalent Number of Parameters: 4.77 Residual Standard Error: 8.608 Trace of smoother matrix: 5.27 (exact) Control settings: span : 0.75 degree : 2 family : gaussian surface : interpolate cell = 0.2 normalize : TRUE parametric : FALSE drop.square: FALSE > plot(x,y)
Output
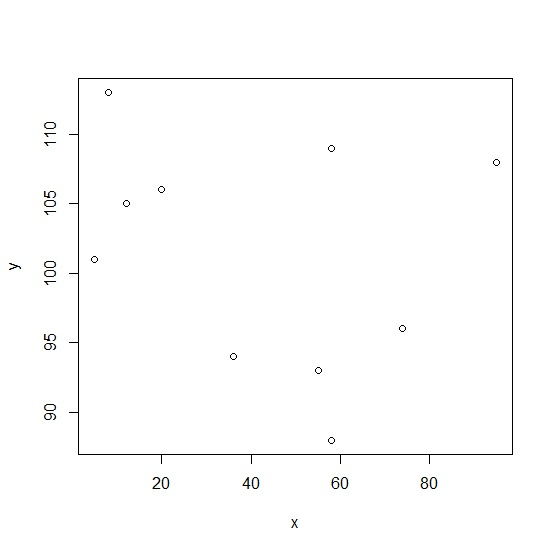
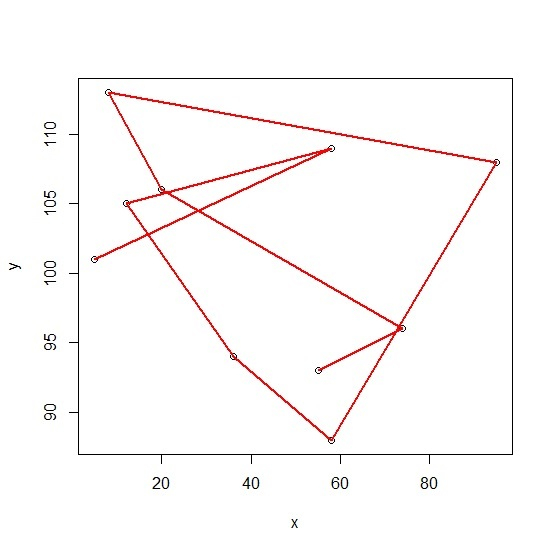
> lines(Model, col='red', lwd=2)
Output

Advertisements
