Article Categories
- All Categories
-
 Data Structure
Data Structure
-
 Networking
Networking
-
 RDBMS
RDBMS
-
 Operating System
Operating System
-
 Java
Java
-
 MS Excel
MS Excel
-
 iOS
iOS
-
 HTML
HTML
-
 CSS
CSS
-
 Android
Android
-
 Python
Python
-
 C Programming
C Programming
-
 C++
C++
-
 C#
C#
-
 MongoDB
MongoDB
-
 MySQL
MySQL
-
 Javascript
Javascript
-
 PHP
PHP
-
 Economics & Finance
Economics & Finance
What are Retrotransposons and What Is Their Role in Epigenetic Regulation?
Introduction
Transposons are a type of mobile genetic element in which DNA sequences have a unique ability to move from one place to another within a chromosome. Eukaryotes have transposons that are structurally similar to bacterial transposons and some similar transposition mechanisms. In other cases, however, the mechanism of transposition seems to involve an RNA intermediate with the evolution of certain classes of RNA viruses.
Epigenetic regulation can be defined as the phenomenon by which the mode of working of a specific gene is controlled by mea
Retrotransposons
Retrotransposons code for an enzyme that is very similar to the enzyme reverse transcriptase and the coding regions are flanked by long terminal repeat (LTR) sequences. They jump from one position to another in the cellular genome with the help of an RNA intermediate, using the enzyme reverse transcriptase that makes the DNA copy of the RNA which is then followed by the ligation of the DNA to the target site.
This method of transposition is commonly seen in eukaryotes and is quite different from that of bacterial transposition where DNA is directly attached to the target site within the chromosomes or at another chromosome.
Retrotransposons do not contain env gene that codes for viral capsid so they cannot form viral particles. This brings us to the comparison of reverse transcriptase and eukaryotic transposons which suggest that reverse transcriptase was present much before multicellular organisms.
Types of retrotransposons
Based on long terminal repeats there are two types of retrotransposons-
LTR retrotransposons
They contain a central coding region that is flanked by LTR that are lying in the same direction, this forms the basic structure of all LTR retrotransposons.
They contain short, inverted repeats on either side in their few hundred nucleotide pair long sequence just like other transposons.
They are named as LTR transposons because of the LTR sequence and properties similar to the retrovirus.
They utilize reverse transcriptase and mediate synthesis of double-stranded DNA from RNA which is then transposed to the target site and ligated to it.
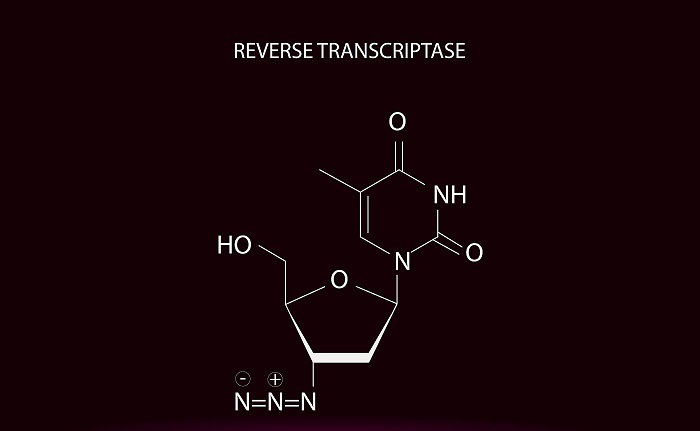
Fig. Reverse transcriptase
The most common type of LTR retrotransposon found in yeast is the Ty element. They are 5.9 kilobase pairs long out of which the length of LTR is about 340 base pairs at both ends. There are about 35 copies of the Ty element found in the yeast. Just like gag and pol genes of retroviruses yeast Ty elements contain TyA and TyB genes.
Non-LTR Retrotransposons
They are the most common type of retrotransposons in eukaryotes. They are also known as retroposons and do not contain long terminal repeats flanking their ends but contain a homogenous sequence of A and T base pairs and one of its terminals which is the modification of poly-A tail.
Process of transposition involves the formation of RNA intermediate which is then converted into double-stranded DNA by reverse transcriptase that is encoded by the element itself and then transposed to the target site.
-
The most common example of these retrotransposons are human transposons namely long interspersed nuclear elements (LINE) and short interspersed nuclear elements (SINE).
Long Interspersed Nuclear Element (LINE)- Human transposable element which belong to this class is L1 retroposon, which is about 6kb long and has internal promoter sequence recognized by RNA polymerase II and two ORFs.
Short Interspersed Nuclear Element (SINE)- They are 400 base pairs long that contain internal promoter sequence but do not code for any proteins. They are the second most abundant transposable element in in humans. They do not contain any terminal repeats but instead have A and T sequences at one end. One of the common examples of SINE element is Alu sequences that are active.
Role of Retrotransposons in Epigenetic Regulation
Transposons make up a large part of human genome, which are capable of jumping from one site to another in chromosome, and sometimes their insertion cause severe damage to a functional gene. Despite having certain functions most of these transposable elements are silenced either by methylation or by histone modification to avoid any kind of deleterious impact on the functional genes.
But sometimes, these elements escape these two processes and cause a change in the function of the gene this process is known as epigenetic evolution. Their escape makes the sites prone to environmental stimuli like exposure to toxins and unhealthy diets by acting as loci of interindividual epigenetic variability.
Example
One of the most common examples of the impact of retrotransposons on epigenetic regulation is the Agouti viable yellow mouse. The retrotransposon found in these mice is Intracisternal A-particle (IAP) which has a deeper impact on DNA methylation that results in interindividual phenotypic variations like coat colors, etc. However, the detailed mechanism is still to be unleashed and studied.
Conclusion
Class I transposable elements are also known as retrotransposons or transposons with RNA intermediates. These elements insert themselves at various genomic loci by converting single-stranded RNA to double-stranded DNA with the help of the enzyme reverse transcriptase. They increase the copy number faster than the DNA transposons.
They are also believed to be involved in the epigenetic regulation of genes and different characteristics imparted by them which was observed in Agouti mice varieties by escaping methylation and histone modification process. Retrotransposons are the most understudied topics of molecular biology; deeper studies can unleash the other impacts of retrotransposons in different organisms.


