Article Categories
- All Categories
-
 Data Structure
Data Structure
-
 Networking
Networking
-
 RDBMS
RDBMS
-
 Operating System
Operating System
-
 Java
Java
-
 MS Excel
MS Excel
-
 iOS
iOS
-
 HTML
HTML
-
 CSS
CSS
-
 Android
Android
-
 Python
Python
-
 C Programming
C Programming
-
 C++
C++
-
 C#
C#
-
 MongoDB
MongoDB
-
 MySQL
MySQL
-
 Javascript
Javascript
-
 PHP
PHP
-
 Economics & Finance
Economics & Finance
Selected Reading
How to save an xtable file locally using R?
To save an xtable file locally, obviously we first need to create the xtable and then use the print function for saving the file. Therefore, we require xtable package loaded in R environment and the data set that we want to save as an xtable file. In the below example, we have used iris data in base R for this purpose. Take a look at the example to understand how it works.
Loading xtable package −
library(xtable)
Considering iris data −
Example
data(iris) head(iris,20)
Output
Sepal.Length Sepal.Width Petal.Length Petal.Width Species 1 5.1 3.5 1.4 0.2 setosa 2 4.9 3.0 1.4 0.2 setosa 3 4.7 3.2 1.3 0.2 setosa 4 4.6 3.1 1.5 0.2 setosa 5 5.0 3.6 1.4 0.2 setosa 6 5.4 3.9 1.7 0.4 setosa 7 4.6 3.4 1.4 0.3 setosa 8 5.0 3.4 1.5 0.2 setosa 9 4.4 2.9 1.4 0.2 setosa 10 4.9 3.1 1.5 0.1 setosa 11 5.4 3.7 1.5 0.2 setosa 12 4.8 3.4 1.6 0.2 setosa 13 4.8 3.0 1.4 0.1 setosa 14 4.3 3.0 1.1 0.1 setosa 15 5.8 4.0 1.2 0.2 setosa 16 5.7 4.4 1.5 0.4 setosa 17 5.4 3.9 1.3 0.4 setosa 18 5.1 3.5 1.4 0.3 setosa 19 5.7 3.8 1.7 0.3 setosa 20 5.1 3.8 1.5 0.3 setosa
Creating an Xtable of partial iris data −
Xtable_iris<-xtable(iris[1:30,])
Saving Xtable_iris locally −
Example
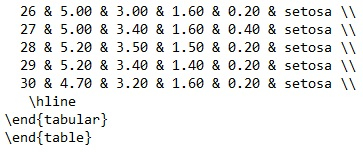
print(Xtable_iris,file="XtableIris30Rows.txt")
Output

Advertisements


