Article Categories
- All Categories
-
 Data Structure
Data Structure
-
 Networking
Networking
-
 RDBMS
RDBMS
-
 Operating System
Operating System
-
 Java
Java
-
 MS Excel
MS Excel
-
 iOS
iOS
-
 HTML
HTML
-
 CSS
CSS
-
 Android
Android
-
 Python
Python
-
 C Programming
C Programming
-
 C++
C++
-
 C#
C#
-
 MongoDB
MongoDB
-
 MySQL
MySQL
-
 Javascript
Javascript
-
 PHP
PHP
-
 Economics & Finance
Economics & Finance
Aminoacyl tRNA synthetase: Mechanism, Structure, and Applications
Introduction
Protein synthesis, the process of creating proteins from amino acids, is a fundamental process in all living organisms. This process requires the cooperation of many different molecules, including ribosomes, transfer RNAs (tRNAs), and aminoacyl tRNA synthetases (aaRSs). In this article, we will focus on aaRSs, the enzymes that attach amino acids to tRNAs, which are essential for the correct reading of the genetic code and the accurate synthesis of proteins.
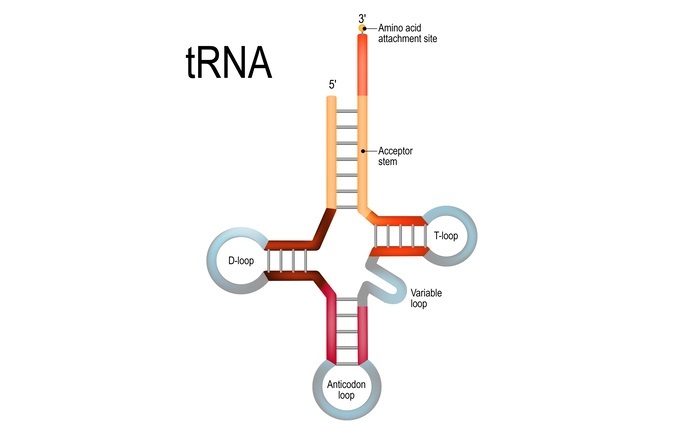
What is Aminoacyl tRNA synthetase?
Aminoacyl tRNA synthetases (aaRSs) are enzymes that catalyze the attachment of specific amino acids to their corresponding tRNAs, a process called aminoacylation or charging.
The aaRSs ensure that each amino acid is attached to the appropriate tRNA, which in turn binds to the mRNA codon that specifies that amino acid.
Thus, aaRSs play a crucial role in maintaining the fidelity of the genetic code, as errors in aminoacylation can lead to the misincorporation of amino acids and ultimately to malfunctioning proteins.
Evolution of Aminoacyl tRNA synthetase
AaRSs are ancient enzymes that have evolved over billions of years to accommodate the diverse amino acid repertoires of different organisms.
The current consensus is that the earliest aaRSs were probably RNA-based enzymes that catalyzed the attachment of amino acids to their corresponding tRNAs in the absence of protein enzymes.
Later, as proteins evolved, some of these RNA-based aaRSs may have been replaced by protein-based ones, which offered greater efficiency and specificity.
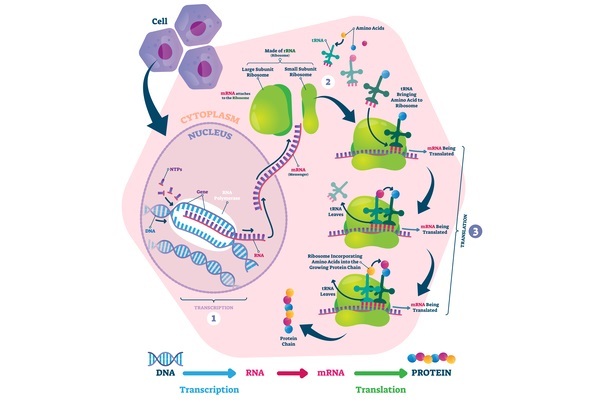
Types of Aminoacyl tRNA synthetase
There are two major classes of aaRSs: class I and class II, which differ in their structural and catalytic features.
Class I aaRSs generally attach amino acids with hydrophobic side chains, such as leucine and isoleucine
While class II aaRSs attach amino acids with polar or charged side chains, such as glutamate and lysine.
Within each class, there are further subdivisions based on the specific amino acids and tRNAs that are recognized.
Mechanism of Aminoacyl tRNA Synthetase
The aminoacylation reaction catalyzed by aaRSs occurs in two steps: activation and transfer. In the activation step, the aaRSs use ATP to activate the amino acid, forming an aminoacyl-AMP intermediate.
In the transfer step, the activated amino acid is transferred from the aaRS to the tRNA, forming an aminoacyl-tRNA complex. The transfer step is highly specific, as aaRSs recognize both the amino acid and the anticodon loop of the tRNA.
Structure of Aminoacyl tRNA synthetase
The structure of aaRSs is highly conserved across different organisms, despite the diversity of amino acid sequences and catalytic mechanisms.
Class I aaRSs are typically composed of two globular domains, one of which binds the amino acid and the other of which binds the tRNA.
Class II aaRSs, on the other hand, typically have a single catalytic domain that binds both the amino acid and the tRNA.
The active sites of aaRSs are often located in clefts or grooves between the domains, which facilitate the recognition and binding of the substrates.
Applications of Aminoacyl tRNA synthetase
AaRSs have many practical applications in both basic research and biotechnology.
One important application is in the creation of artificial genetic systems that incorporate unnatural amino acids into proteins.
By engineering aaRSs to recognize and attach new amino acids, researchers have been able to expand the genetic code and create novel proteins with new properties and functions.
Another application of aaRSs is in the development of antibiotics that target bacterial aaRSs, inhibiting protein synthesis and killing the bacteria.
Finally, aaRSs are also used in the production of synthetic peptides and proteins for therapeutic and diagnostic purposes.
Conclusion
In conclusion, aminoacyl tRNA synthetases are key enzymes in protein synthesis that ensure the accurate attachment of amino acids to tRNAs. Through their evolution and diverse structures, aaRSs have enabled the genetic code to be read and translated into functional proteins in all living organisms. Their applications in research and biotechnology highlight the importance of understanding these enzymes and their mechanisms in order to manipulate and harness their potential.
FAQs
Q1. How many aminoacyl tRNA synthetases are there in humans?
Ans. There are 20 different aaRSs in humans, each specific for one of the 20 amino acids.
Q2. Can aminoacyl tRNA synthetases make mistakes?
Ans. Yes, aaRSs can make mistakes and attach the wrong amino acid to a tRNA, leading to the misincorporation of amino acids and potentially harmful effects.
Q3. How do antibiotics that target aaRSs work?
Ans. Antibiotics such as tetracycline and macrolides bind to bacterial aaRSs and inhibit their activity, preventing the attachment of amino acids to tRNAs and blocking protein synthesis.
Q4. Can aaRSs be used to create proteins with new functions?
Ans. Yes, by engineering aaRSs to recognize and attach new amino acids, researchers can create novel proteins with new properties and functions.
Q5. Are aaRSs essential for all living organisms?
Ans. Yes, aaRSs are essential for all organisms that use the genetic code to synthesize proteins.


